Applying Breakthroughs in Protein Structure Calculation to the Creation of Designer Enzymes
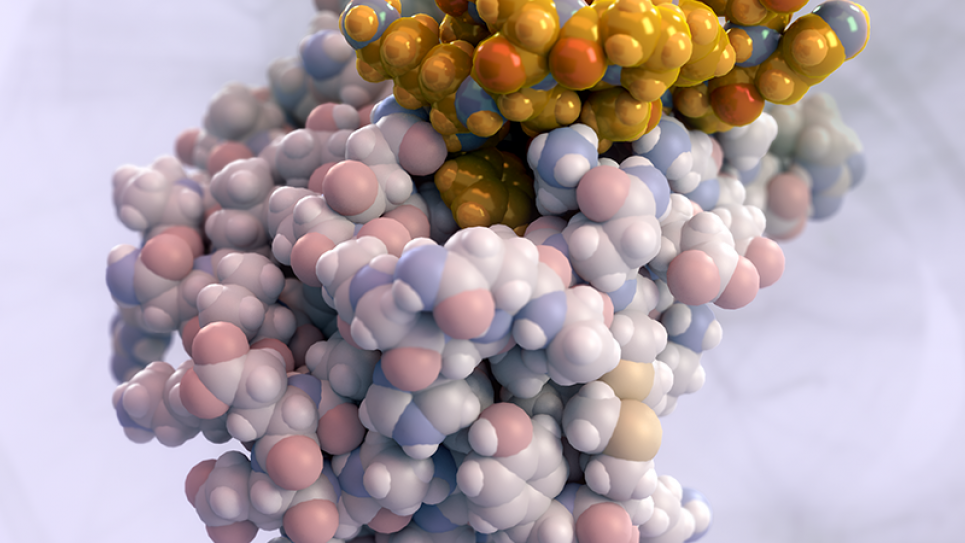
Proteins are large molecules found in all living organisms that serve as the major functional units of all life. Proteins are made from a sequence of amino acid building blocks. Key to understanding proteins is to understand the relationship between the amino acid sequence, the three‐dimensional folded structure, and the resulting protein function. The Baker laboratory has developed the Rosetta software suite for predicting protein structure given the amino acid sequence, and for designing amino acid sequences capable of folding into novel structures with desired functions. Of particular interest are enzymes, a type of protein responsible for catalyzing (aiding) chemical reactions. Enzymes serve as nature’s biomachinery and are responsible for invaluable functions such as converting food to energy, building cell walls, and repairing DNA. Recent advances in massively parallel design algorithms have opened the door for the design of artificial enzymes capable of catalyzing chemical reactions that no natural enzyme can catalyze. Designer enzymes could have broad applicability in many DOE priority areas, such as sustainable domestic biofuel production, and could reduce energy consumption and improve energy efficiency by permitting industrial syntheses to occur at lower temperatures and with fewer unwanted byproducts. We propose to use a multistate protein design strategy dependent on large-scale parallel supercomputing hardware to create designer enzymes with a broad range of applications.