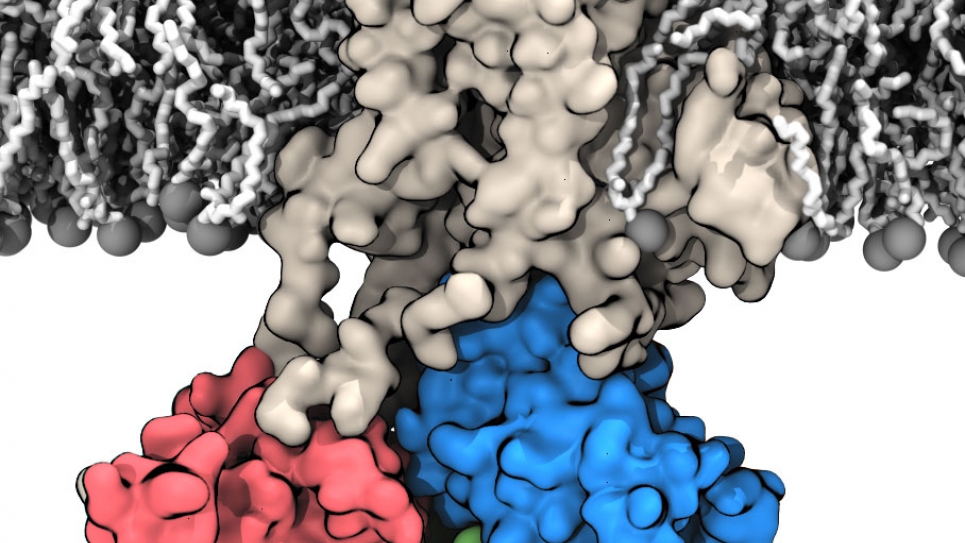
The Free Energy Landscapes Governing Membrane Protein Function
Large membrane proteins involved in transport and cellular signaling are critical for life. The objective of this research is to elucidate the mechanism that underlies their function at the molecular level. This proposal is focused on three prototypical important membrane proteins: the ATP-driven sarcoplasmic endoplasmic reticulum calcium pump SERCA, the voltage-activated potassium channel, and the voltage-activated phosphatase CiVSP. A complete understanding of how these proteins carry out their function will rely heavily on characterizing conformational transitions. However, such conformational transition pathways, and their associated free energies, are challenging to determine directly from experiments due to their transient, short-lived states.
To determine functionally important conformational transition pathways and characterize the energetics associated with these structural changes, researchers will utilize the unifying concept of free energy landscape. Considerable progress has already been made in discovering the transition pathways of these systems using the string method simulations. However, a more complete understanding of their mechanism requires a quantitative characterization of the pathways and free energy landscapes that govern motions along them. This vision can be achieved by leveraging the information from classical molecular dynamics simulations of atomic-resolution models.
By studying three prototypical membrane proteins of increasing size and complexity within a unified theoretical framework based on free energy landscapes, this project will push the envelope and advance the theory-modeling-simulation (TMS) technology. TMS offers a virtual route to address fundamental biological questions and help solve the problem of rational protein design. The study will serve as a road-map for simulating, visualizing, and elucidating how biomolecular nano-machine membrane proteins work.