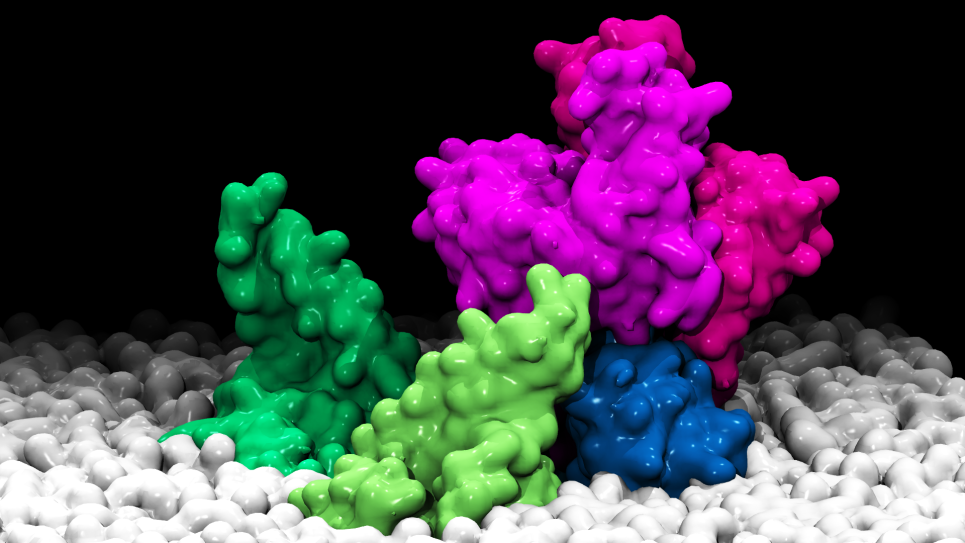
A living cell is made of many different smaller pieces including macromolecules called proteins. Proteins are built from atoms and many work as nanoparticle-sized biomachinery for the cell. Living cells can be viewed as a collection of these proteins and other macromolecules carrying out their well-defined tasks in the cellular environment. In order to perform their tasks, proteins often come together and form a complex. The ability to recognize their partner and form a tight complex is important. Disruption of complex formation due to mutation, often leads to loss of function and can lead to disease, such as cancer. Scientists therefore need to gain a deep understanding on protein recognition and assemblies at the atomic level.
To achieve the goal of understanding protein recognition and complex formation, this research team has developed powerful and scalable computational methodologies that are well adapted for leadership computers. The team’s method breaks-down the binding process into several physically meaningful steps and expresses the total binding free energy – the energy associated with complex-formation – as a sum of free energies associated with each of these smaller steps. This step-by-step framework provides a rational and tractable starting point to characterize protein-protein interaction quantitatively and explain how mutations and structural changes are associated with observed thermodynamic effects.
This allocation supports rigorous testing of atomic scale (molecular dynamics) free energy simulation methods by considering a set of representative protein complexes of increasing size and complexity. The results will support determination of best practices and the building of a pipeline that automates the simulation setup according to the chosen protocol. This computational infrastructure will reduce human error and rapid simulation setup/execution in high performance computers. The research will result in great technical advances with cutting-edge computational methodologies, and generate new knowledge about an important class of biological systems.