Optimizing a deep learning framework for cancer research for Aurora
As part of a series aimed at sharing best practices in preparing applications for Aurora, we highlight researchers' efforts to optimize codes to run efficiently on graphics processing units.
Leveraging supercomputers for cancer research
Developed as part of the Exascale Deep Learning and Simulation Enabled Precision Medicine for Cancer project, CANDLE (or the CANcer Distributed Learning Environment), is an exascale-optimized framework that seeks to leverage the next generation of supercomputing to answer broad questions cancer researchers face.
Best practice
- Use standard profiling tools such as Intel VTune and NVIDIA Nvprof or NSight to pinpoint sections of code that present potential bottlenecks.
CANDLE will be deployed on U.S. Department of Energy (DOE) supercomputers such as the forthcoming Aurora system, an exascale Intel-HPE machine to be housed at the Argonne Leadership Computing Facility (ALCF), a U.S. DOE Office of Science User Facility located at Argonne National Laboratory.
The software suite, which is supported by DOE's Exascale Computing Project and ALCF Early Science Program, broadly consists of three components: a collection of deep neural networks that capture and represent different problems in cancer, a library of code adapted for exascale-level computing and a component that orchestrates how work will be distributed across the computing system.
Lesson learned
- Machine learning and deep learning-based applications such as CANDLE can only be successfully ported subsequent to the successful implementation of their underlying software framework.
Solving problems at multiple scales
The project seeks to implement deep learning architectures both relevant to cancer research and capable of addressing critical problems therein at three biological scales: Pilot 1 (P1), cellular; Pilot 2 (P2), molecular; and Pilot 3 (P3), population.
- P1 benchmarks, arising from cellular-level problems and data, at a high level aim to predict drug responses based on the particular molecular qualities of tumor cells and drug descriptors.
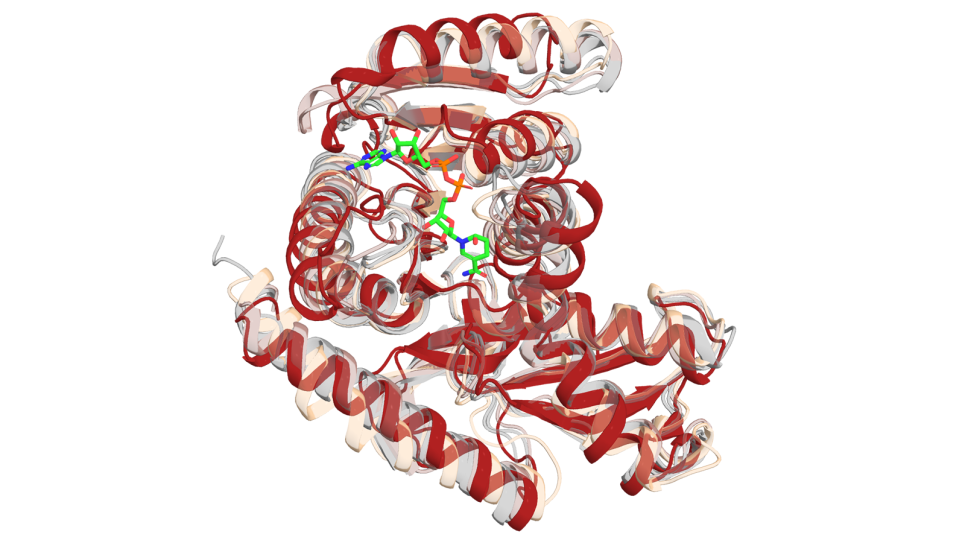
- P2 benchmarks deal with problems in molecular dynamic simulations of proteins involved in cancer. The specific proteins are known as RAS proteins, a family of protein encoded within the Ras gene that can become oncogenic when mutated.
- P3 benchmarks, generated from population-level problems and data, are used to predict cancer recurrence in patients based on patient-related information.
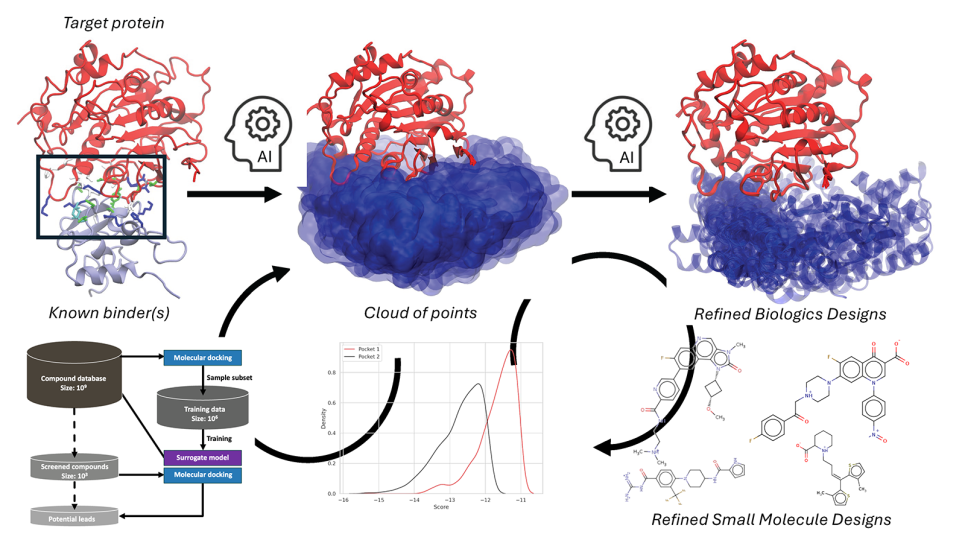
Here we focus on one application Uno, a deep learning model CANDLE applies to P1 benchmarks, which aims to predict tumor response to both single and paired combinations of drugs based on the molecular features of tumor cells assessed across multiple data sources. The publicly available code implements a deep learning architecture with 21 million parameters in the TensorFlow framework and, already running at scale on DOE supercomputers such as Theta and Summit, is being prepared for Aurora.
As the massive growth in the number and variety of drugs to be screened and simulated can tax traditional hardware to its limits, the increase in computational power expected of exascale promises to greatly benefit and accelerate the design of effective drugs and drug combinations.
Porting to exascale
Efforts to port CANDLE Uno to exascale began with the developers working to attain a deep understanding of the application’s performance on existing hardware such as Intel KNLs and NVIDIA GPUs. The developers evaluated the efficiency of the application performance using throughput as a metric—that is, they measured the number of samples processed per second.
Such evaluation included profiling the code with standard performance analysis tools, such as using Intel VTune and NVIDIA Nvprof to pinpoint any potential bottlenecks in the code that could hinder execution of the application when run at scale. To enable execution on Aurora, Intel Data Parallel C++ (DPC++) is utilized within the Intel implementation of the deep learning software TensorFlow and PyTorch. This setup encourages effective performance from the models Intel GPUs.
Bottlenecks are addressed through incremental releases of deep learning software for exascale. For instance, after observing low GPU utilization, the developers used profiling tools to identify the cause as diminished kernel size, a problem subsequently tackled by improving the manner in which the Intel deep learning framework optimizes the graph implementation. This also resulted in improved application throughput.
Finally, the developers tested the code incorporating incremental releases of TensorFlow developed by Intel for ever-improving GPU-optimization.
While the application is hardware-agnostic, its particular implementation relies on the deep learning framework. This means that while the code will run on Aurora and other DOE supercomputers alike, its implementations may vary if two given machines incorporate different frameworks. Differences in the underlying hardware (for example, an Intel GPU versus an NVIDIA GPU), on the other hand, may lead to the presence of various custom-tuning techniques.
As facilitated by TensorFlow, the code is being developed in such a way that compute-heavy tasks are delegated to GPUs whenever they are available for processing. Deep learning models, for instance, are trained in GPU runs, whereas tasks such as data loading are consigned to CPUs.
Apart from Uno, similar efforts to port other applications from CANDLE to run on Intel GPUs are ongoing.
Successful framework implementation
Machine learning and deep learning-based applications such as CANDLE can only be successfully ported subsequent to the successful implementation of their underlying software framework—an ecosystem like Python, TensorFlow, PyTorch, or Keras. Successful implementation occurs in a two-step process.
First the functionality of the application must be tested—that is, the code must successfully execute in full in a limited setting without incurring any serious bugs. The software framework implementation requires further optimization to help the application satisfy the convergence criteria with maximum performance metrics, such as throughput.